  
This mutant reproducibly showed significantly less FI* formation than the wild-type template. The formation of FI* was observed for mutant oriV 3–6, however, densitometric quantitation indicated that only 33% of the original supercoiled template was converted to the FI* form. The lack of DnaB unwinding activity in mutant oriV 2–6 suggests that the small amount of strand opening that was observed in oriV 2–6 (see Fig. No FI* was observed for mutant templates oriV 1–6 and oriV 2–6. Upon the addition of His6-TrfA 254D/267L protein to the reaction, 50% (using densitrometry scans) of the wild-type oriV template was converted to the faster migrating FI* species (Fig. The same assay was used to compare wild-type and mutant oriVtemplates for their ability to serve as substrates for DnaB helicase unwinding. Mutant oriV 3–6 displayed the same pattern of modification as wild-type oriV on both stands, indicating that this insertion upstream of the iterons had no affect on the formation of the TrfA♽naA-mediated open complex. Thus, the insertion in the M2 13-mer disrupted the region of helical instability required for strand opening. Similarly, the bottom strand analysis of oriV 2–6 showed a reduced level of modification with a shift in some of the modified thymines at the beginning of the M2 13-mer (Fig. However, no modification of the middle 13-mers at the site of the 6-bp insertion was observed. Mutant oriV 2–6 exhibited some modification at the positions of the L and R 13-mers although the pattern of modification at the L 13-mer was shifted upward in the gel. Mutant oriV 1–6 which has a 6-bp insertion between the iterons and the 13-mers exhibited no modification by KMnO 4 in the 13-mer region on either strand indicating the absence of any open structure. coli strain, CC118, where replication can be initiated only from the RK2 origin. To test for oriV function in vivo, plasmids were purified through two CsCl buoyant density gradients and between 200 and 500 ng was used for the transformation of the parent E. coli strain CC118 (λ pir) on Luria-Bertani agar plates supplemented with 12.5 μg/ml tetracycline. 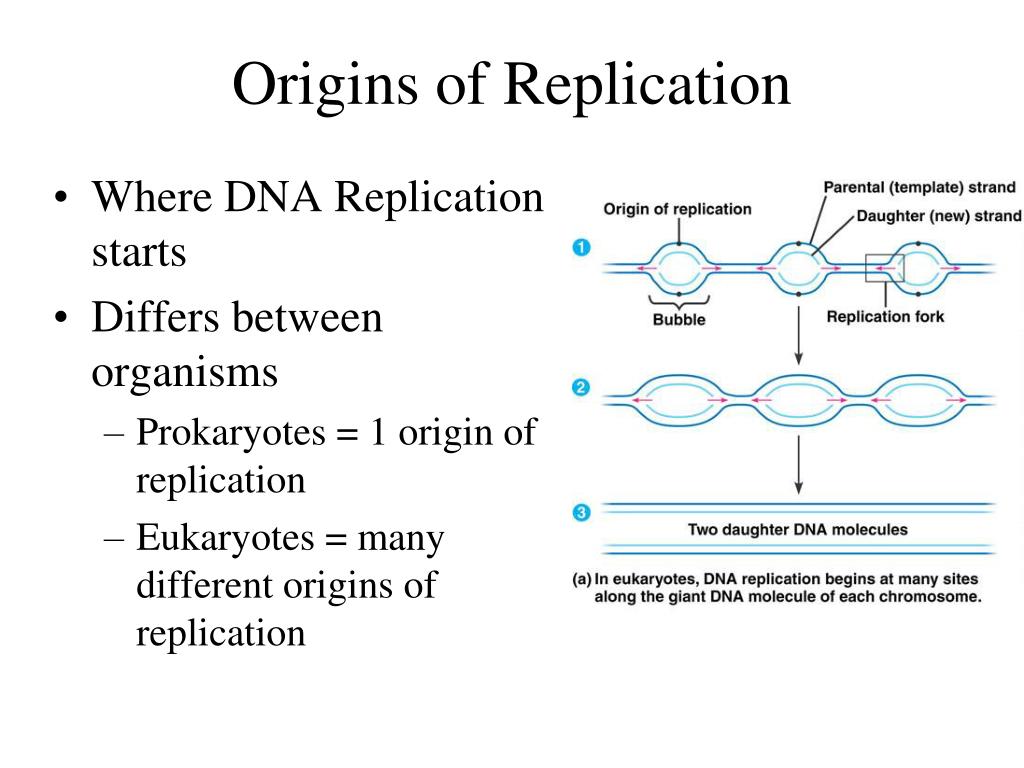
Plasmids containing a mutated oriV in pKD19L1 were obtained and maintained in E. Unlike the other derivatives, oriV Δ GC was not generated by PCR but by cutting pKD19L1 oriV 4–6 (which has an additional BamHI site due to the insertion) with BamHI and religating which resulted in a deletion of 65 bp. the KpnI- XbaI fragment from pSP6 containing the mutated oriV sequence was subcloned into pKD19L1 replacing the wild-type origin. Amplification on template DNA, pSP6, was for 9–18 cycles at 95 ☌ for 1 min, 56 ☌ for 1 min, and 68 ☌ for 8 min. Primers used for mutagenesis were designed according to manufacturers recommendations and included the recognition sequence for either EcoRI or BamHI. These results demonstrate that correct helical phasing and intrinsic DNA curvature are critical for the formation of an open complex and that the DnaA boxes must be on the correct side of the helix to load DnaB helicase. In these mutants, however, strand opening appeared normal but the levels of DnaB helicase activity were substantially reduced. Insertions of more or less than one helical turn between the DnaA boxes and the iterons also inactivated the replication origin. Mutations in the region between the iterons and the 13-mers which altered the helical phase or the intrinsic DNA curvature prevented strand opening of the origin and consequently abolished replication activity. These altered origins were analyzed for replication activity in vivo and in vitro, for localized strand opening and for DnaB helicase mediated unwinding. To investigate how the organization of the RK2 origin contributes to the mechanism of replication initiation, mutations were introduced into the minimal origin which altered the sequence and/or spacing of each particular region relative to the rest of the origin. The 393-base pair minimal origin, oriV, of plasmid RK2 contains three iterated motifs essential for initiation of replication: consensus sequences for binding the bacterial DnaA protein, DnaA boxes, which have recently been shown to bind the DnaA protein 17-base pair direct repeats, iterons, which bind the plasmid encoded replication protein, TrfA and A + T-rich repeated sequences, 13-mers, which serve as the initial site of helix destabilization. Glycobiology and Extracellular Matrices.Replication of DNA in prokaryotes begins at a single origin of replication, shown in the figure to the left, and proceeds in a bidirectional manner around the circular chromosome until replication is complete. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
